|
Haptics and its Applications to Medicine
Ingrid Carlbom, Stefan Seipel, Pontus Olsson, Fredrik Nysjö
Partner: Stefan Johansson (Division of Microsystems Technology, UU and Teknovest AB); Jan-Michaél Hirsch, Dept. of Surgical Sciences, Oral & Maxillofacial Surgery, UU and Consultant at Dept. of Plastic- and Maxillofacial Surgery, UU Hospital; Andreas Thor, Dept. of Surgical Sciences, Oral & Maxillofacial Surgery, UU Hospital; Andres Rodriguez Lorenzo, Department of Surgical Sciences, Plastic Surgery, UU Hospital; PiezoMotors AB, SenseGraphics AB.
Funding: Dept. of Surgical Sciences, Oral & Maxillofacial Surgery, University Hospital
Period: 1301-
Abstract:
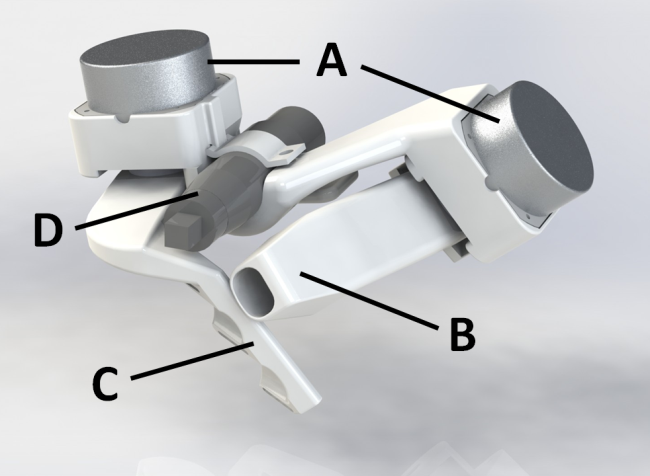
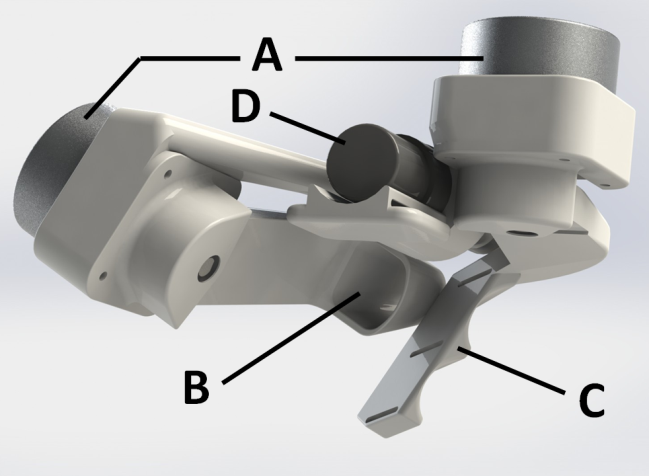
Two Degrees-of-Freedom Haptic Gripper with Ultrasonic Piezoelectric Motors Piezoelectric motors have a high force/mass ratio, which makes them a promising alternative to electromagnetic motors for actuation of haptic interfaces. We have previously developed and evaluated a haptic gripper actuated by a quasi-static piezoelectric motor, operating within the audible range. The evaluation highlighted two main areas for improvement: faster and quieter actuation. During the last year we have designed a new admittance-type haptic gripper (see Figure 18) with two degrees-of-freedom (DOF), actuated by ultrasonic piezoelectric motors with higher maximum speed and silent operation compared to quasi-static motors. The gripper provides one DOF for the thumb and one DOF for the remaining fingers. All DOFs are direct-drive, involving no mechanical gearing, to minimize backlash and friction. Two custom-made strain-gauge load cells, mounted on the motor axes, measure forces applied by the user.


|
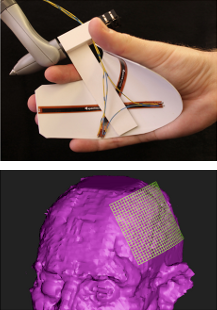
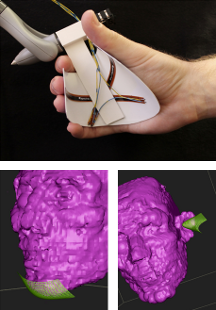
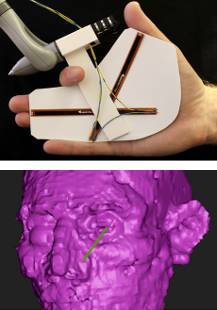
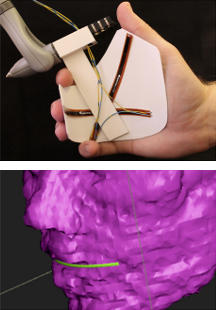
SplineGrip - An Eight Degrees-of-Freedom Flexible Haptic Sculpting Tool SplineGrip is a flexible haptic sculpting tool that senses the articulation of the hand in two degrees-of-freedom (DOF) that we presented as a SIGGRAPH'13 poster. The tool is mounted on a commercial haptic device that tracks hand pose (position and orientation in six DOF) while simultaneously providing three DOF haptic feedback to the hand. The eight DOF input is mapped to the pose and shape of a virtual representation of a sculpting tool (Figure 19), offering versatile interaction with a virtual model.


|
|
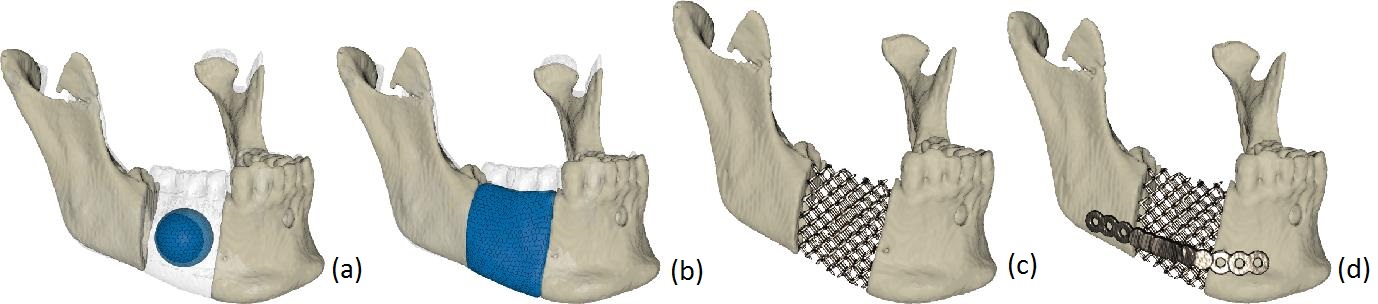
We developed a semi-automatic method based on deformable models and haptics that allows the design and virtual testing of a scaffold implant before production. Using pre-segmented CT-data, a surgeon begins with virtual bone resection, using an interactive cutting tool, to give the defect region good load bearing contact surfaces. Next he/she positions the initial deformable model, which may be a simple sphere, in the defect region and places the bounding surface around the defect (Figure 20a). The system calculates external forces for the deformable model that expand it towards the defect contact surfaces and the bounding surface until it fills the defect (Figure 20b). The shape may be adjusted interactively. The surgeon may also refine the implant with the cutting tool and reposition it inside the defect relying on haptic feedback to perfect its fit. The system generates the scaffold structures of the implant (Figure 20c), and the surgeon may add fixation plates to the structure (Figure 20d).
ProViz - Interactive Visualization of 3D Protein Images
Lennart Svensson, Ida-Maria Sintorn, Ingela Nyström, Fredrik Nysjö, Johan Nysjö, Anders Brun, Gunilla Borgefors
Partners: Dept. of Cell and Molecular Biology, Karolinska Institute; SenseGraphics AB
Funding: The Visualization Program by Knowledge Foundation; Vaardal Foundation; Foundation for Strategic Research; VINNOVA; Invest in Sweden Agency
Period: 0807-
Abstract: Electron tomography is the only microscopy technique that allows 3-D imaging of biological samples at nano-meter resolution. It thus enables studies of both the dynamics of proteins and individual macromolecular structures in tissue. However, the electron tomography images have a low signal-to-noise ratio, which makes image analysis methods an important tool in interpreting the images. The ProViz project aims at developing visualization and analysis methods in this area.
In general, the project focus 2013 has been on increasing the accessibility of the previously developed methods, by continuing to work on a user-friendly software containing the most important ProViz results. Figure 21 shows how this software can display an electron tomogram, synthetic in this case, and a 3-D fitness landscape showing the matching results for a protein template.
Project highlights during 2013 include a two months research collaboration stay at the Okinawa Institute of Science and Technology, OIST, in Japan and the publishing of a paper at the Iberian Conference on Pattern Recognition and Image Analysis, IbPRIA. During the stay at OIST the software developed in the ProViz project was presented and discussed in depth, with adjustments to the software and improvements to an upcoming manuscript as the result. The research stay was made possible primarily through a JSPS, Japan Society for the Promotion of Science, fellowship. The presented paper concerned a new way of creating registration templates for finding biological structures in electron tomograms.
|
|
Analysis and Processing of Three-Dimensional Magnetic Resonance Images on Optimal Lattices
Elisabeth Linnér, Robin Strand
Funding: TN-faculty, UU
Period: 1005-
Abstract: Three-dimensional images are widely used in, for example, health care. With optimal sampling lattices, the amount of data can be reduced by 30% without affecting the image quality. In this project, methods for image acquisition, analysis and visualization using optimal sampling lattices are studied and developed, with special focus on magnetic resonance imaging. The intention is that this project will lead to faster and better processing of images with less demands on data storage capacity.
During 2013, the focus has been on distance transforms. A paper describing a graph-based implementation of the anti-aliased Euclidean distance transform was submitted for publication.
Registration of Medical Volume Images
Robin Strand, Filip Malmberg
Partner: Joel Kullberg, Håkan Ahlström, Dept. of Radiology, Oncology and Radiation Science, UU
Funding: Faculty of Medicine, UU
Period: 1208-
Abstract: In this project, we mainly process magnetic resonance tomography (MR) images. MR images are very useful in clinical use and in medical research, e.g., for analyzing the composition of the human body. At the division of Radiology, UU, a huge amount of MR data, including whole body MR images is acquired for research on the connection between the composition of the human body and disease.
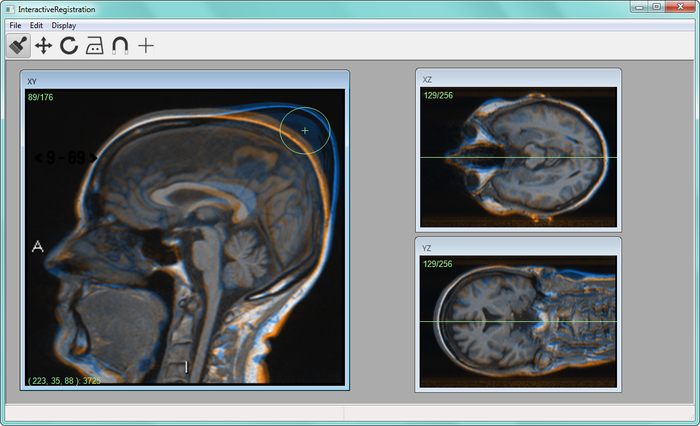
To compare volume images voxel by voxel, we develop image registration methods. For example, large scale analysis is enabled by image registration methods that utilizes, for example, segmented tissue (see, e.g., Project 34) and anatomical landmarks. Another example is interactive image registration where a user can fine-tune the segmentation result by a user interface, see Figure 22.
|
Interactive Segmentation and Analysis of Medical Images
Filip Malmberg, Robin Strand, Ingela Nyström, Ewert Bengtsson
Partners: Joel Kullberg, Håkan Ahlström, Dept. of Radiology, Oncology and Radiation Science, UU
Funding: TN-faculty, UU
Period: 1106-
Abstract: Three-dimensional imaging technique such as computed tomography (CT) and magnetic resonance imaging (MRI ) are now routinely used in medicine. This has lead to an ever increasing flow of high-resolution, high-dimensional, image data that needs to be qualitatively and quantitatively analyzed. Typically, this analysis requires accurate segmentation of the image.
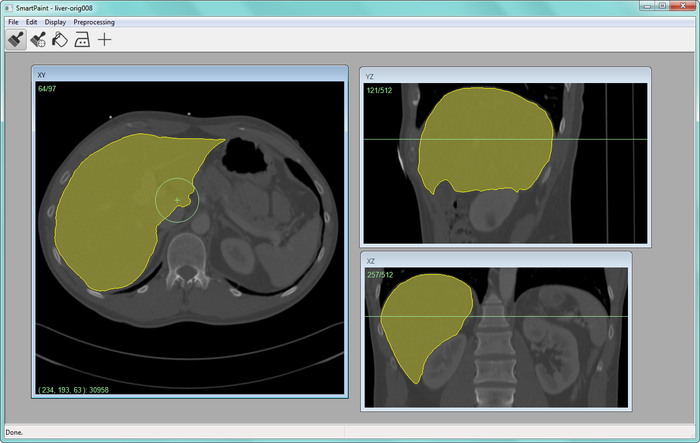
At CBA, we have been developing powerful new methods for interactive image segmentation (see Project 39). In this project, We seek to employ these methods for segmentation of medical images, in collaboration with the Dept. of Radiology, Oncology and Radiation Science at the UU Hospital. In 2013 a software for interactive segmentation, called Smartpaint, was made publicly available (Fig. 23). The software can be downloaded from http://www.cb.uu.se/~filip/SmartPaint/.
|
Orbit Segmentation for Cranio-Maxillofacial Surgery Planning
Filip Malmberg, Ewert Bengtsson, Ingela Nyström, Johan Nysjö
Partners: Jan Michael Hirsch, Andreas Thor, Johanna Nilsson, Dept. of Surgical Sciences, UU Hospital; Roman Khonsari, Pitie Salpetriere Hospital, Paris, France; Jonathan Britto, Great Ormond Street Hospital, London, United Kingdom
Funding: TN-faculty, UU; NovaMedTech
Period: 0912-
Abstract: A central problem in cranio-maxillofacial (CMF) surgery is to restore the normal anatomy of the skeleton after defects, i.e., malformations, tumors and trauma to the face. This is particularly difficult when a fracture causes vital anatomical structures such as the bone segments to be displaced significantly from their proper position, when bone segments are missing, or when a bone segment is located in such a position that any attempt to restore it into its original position poses considerable risk for causing further damage to vital anatomical structures such as the eye or the central nervous system. There is ample evidence that careful pre-operative planning can significantly improve the precision and predictability and reduce morbidity of the craniofacial reconstruction. In addition, time in the operating room can be reduced. An important component in surgery planning is to be able to accurately measure the extent of certain anatomical structures. Of particular interest in CMF surgery are the shape and volume of the orbits (eye sockets) comparing the left side with the right side. These properties can be measured in CT images of the skull, but this requires accurate segmentation of the orbits. Today, segmentation is usually performed by manual tracing of the orbit in a large number of slices of the CT image. This task is very time-consuming, and sensitive to operator errors. Semi-automatic segmentation methods could reduce the required operator time significantly. In this project, we are developing a prototype of a semi-automatic system for segmenting the orbit in CT images. The segmentation system is based on WISH, a software package for interactive visualization and segmentation that has been developed at CBA since 2003. WISH has been released under an open-source license and is available for download at http://www.cb.uu.se/research/haptics.
In 2011, a paper about the orbit segmentation system was presented at the International Visual Information Conference (IVIC) in Malaysia. We also started investigating other applications for the system, e.g., volumetric measurements of the airway space in cone beam CT images and volumetric measurements of the maxillary sinuses in CT images.
|
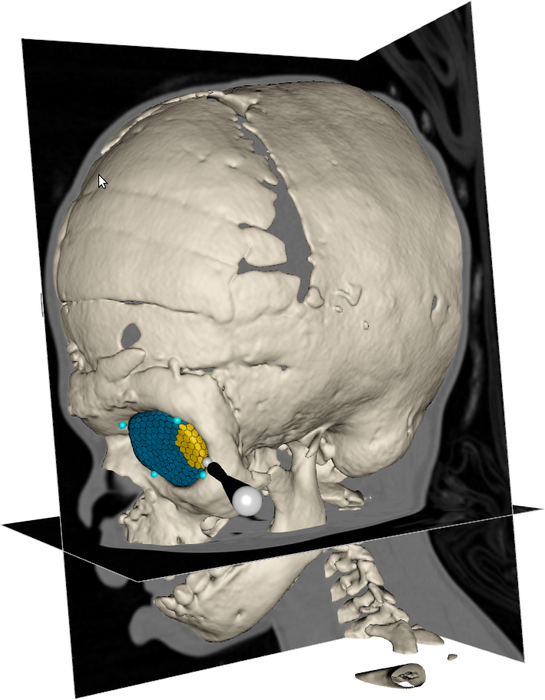
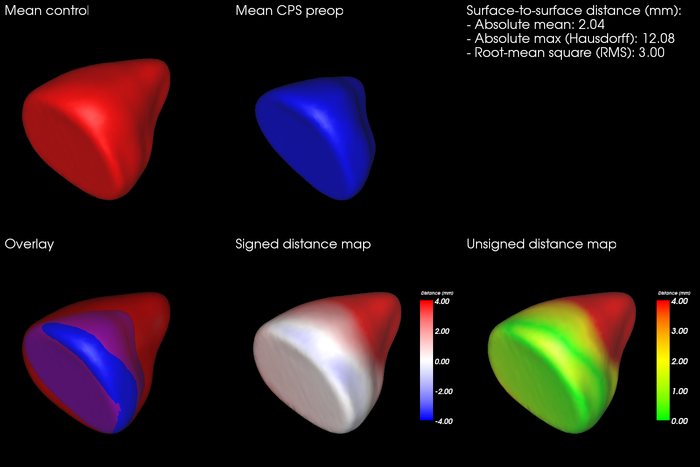
During 2013, we have been collaborating with people from the Craniofacial Centre at Great Ormond Street Hospital, London, United Kingdom, in a project that aims to analyse the size and shape of the orbits in pre- and post-operative CT images of patients with congenital disorders. The semi-automatic segmentation system has been used to segment the orbits in these datasets (Fig. 24), and we have developed automatic tools for performing size and shape analysis of the segmented orbits (Fig. 25). Several abstracts about the ongoing work has been presented at medical conferences. Next, we plan to summarize the collected data and extend the abstracts to journal publications. In addition, we have performed two smaller orbit segmentation studies together with people from the UU hospital, resulting in one presented abstract at Tandläkarnas Riksstämma. In collaboration with Roman Khonsari at Pitie Salpetriere Hospital, Paris, France, we also published a paper on shape and volume measurements on intentionally deformed skulls in American Journal of Physical Anthropology.
|
Precise 3D Angle Measurements in CT Wrist Images
Filip Malmberg, Johan Nysjö, Ingela Nyström, Ida-Maria Sintorn
Partners: Albert Christersson, Dept. of Orthopedics, UU Hospital
Funding: TN-faculty, UU
Period: 1111-
Abstract: To be able to decide the correct treatment of a fracture, for example, whether a fracture needs to be operated on or not, orthopedic surgeons need to assess the details about the fracture. One of the most important factors is the fracture displacement, particularly the angulation of the fracture. When a fracture is located close to a joint, for example, in the wrist, which is the most common location for fractures in the human being, the angulation of the joint line in relation to the long axis of the long bone needs to be measured (Fig. 26a). Since the surface of the joint line in the wrist is highly irregular, and since it is difficult to take X-rays of the wrist in exactly the same projections from time to time, conventional X-ray is not an optimal method for this purpose. In most clinical cases, conventional 2D angle measurements in X-ray images are satisfactory for making correct decisions about treatment, but when comparing two different methods of treatment, for instance, two different operation techniques, the accuracy and precision of the angle measurements need to be higher.
In this project, we are developing a system for performing precise angle measurements in 3D computed tomography (CT) images of the wrist (Fig. 26b). Our proposed system is semi-automatic; the user is required to guide the system by indicating the approximate position and orientation of various parts of the radius bone. This information is subsequently used as input to an automatic algorithm that makes precise angle measurements. A RANSAC-based method for estimating the long axis of the radius bone was presented at the International Conference on Computer Vision and Graphics (ICCVG' 2012). During 2013, we developed a registration-based method for measuring the orientation of the joint surface of the radius. This method was combined with the previously developed axis estimation method and presented at ICIAP 2013. Currently, we are performing a more extensive case study (involving 40 CT scan sequences of fractures wrists) to further evaluate the performance of the 3D angle measurement method and compare it with the conventional 2D X-ray measurement method.
|
Efficient Algorithms for Computer Graphics
Ewert Bengtsson, Anders Hast
Partner: Tony Barrera, Uppsala
Funding: TN-faculty, UU
Period: 9911-
Abstract: Computer graphics is increasingly being used to create realistic images of 3D objects for applications in entertainment, (animated films, games), commerce (showing 3D images of products on the web), industrial design and medicine. For the images to look realistic high quality shading and surface texture and topology rendering is necessary. A proper understanding of the mathematics behind the algorithms can make a big difference in rendering quality as well as speed. We have in this project over the years re-examined several of the established algorithms and found new mathematical ways of simplifying the expressions and increasing the implementation speeds without sacrificing image quality. We have also invented a number of completely new algorithms. The project is carried out in close collaboration with Tony Barrera, an autodidact mathematician. It has been running since 1999 and resulted in more than 25 international publications and a PhD thesis.
During 2013 a poster was accepted for the ACM Computing Frontiers conference entitled: "An Algorithm for Parallel Calculation of Trigonometric and Exponential Functions".
Ubiquitous Visualization in the Built Environment
Stefan Seipel, Fei Liu
Funding: University of Gävle; TN-faculty, UU
Period: 110801-
Abstract: This research project in ubiquitous visualization deals with mobile visualization of spatial data in indoor and outdoor environments. Several key problems for robust mobile visualization are addressed such as spatial tracking and calibration, image based 2D and 3D registration and efficient graphical representations in mobile user interfaces. During 2013 we have devised a fasade region detection method by analyzing image profiles for repetitive patterns in street view images. These profiles are generated by scanning the hue channel of images along lines constructed with edge line segments and vanishing points. The work has been compiled into a paper titled "Detection of Fasade Regions in Street View Images from Split-and-Merge of Perspective Patches" and submitted to the International Conference on Computing and Computer Vision 2014. Meanwhile, we have also been exploring various image features to describe the detected fasade regions in order to identify which building is presented in a specific image.