Khalid Niazi, Ewert Bengtsson
Partners: Mats Nilsson, Div. of Toxicology, Dept. of Pharmaceutical Biosciences, UU; Prof. Bill Webster, School of Medical Sciences, University of Sydney
Funding: COMSATS IIT, Islamabad
Period: 0711-
Abstract: Embryo cultures of rodents is an established technique for monitoring adverse effects of chemicals on embryonic development. The assessment involves determination of the heart rate of the embryo. We have modeled the movement of the heart as a sinusoid and developed a method to represent the heart motion as a function of time. Analysis of the heart beats were carried out using Emperical Mode Decomposition (EMD).
EMD is a powerful data analysis technique which decomposes complicated signal into intrinsic mode functions (IMF). EMD is usually combined with Hilbert transform to analyze the instantaneous frequency in non-stationary signals. This combination is commonly referred to as Hilbert-Huang Transform (HHT). In this project, we have used EMD with slight modification along with Laplacian Eigenmaps to detect periodic activity in rat embryo's heart. The normal embryo's heart activity can easily be detected by local maxima detection but it becomes a challenging task once the heart activity becomes abnormal.
The motion of red blood cells in a vessel also indicates the effects of chemicals on the embryonic development. Currently, we are exploring different methods to analyse the motion of red blood cells.
Khalid Niazi, Ewert Bengtsson, Ingela Nyström
Partners: Mats Nilsson, Lennart Dencker, Div. of Toxicology, Dept. of Pharmaceutical Biosciences, UU
Funding: COMSATS IIT, Islamabad; Swedish Research Council
Period: 0909-
Abstract: The whole embryo culture assay is endorsed as one of few good in vitro embryotoxicity assays available. The goal of the project is to improve the objectivity and sensitivity in the measurements with the help of image and data analysis methods in the assay using three clinical relevant anti-epileptic drugs as model embryotoxicants.
The scoring methods used for morphology are generally semi-quantitative (categorical, score) and mainly associated with growth rate. Specific malformations are scored as either absent (=0) or present (=1). We propose to complement existing scoring systems that will improve embryotoxicity comparisons between labs and enable the creation of a database with cross-reference ability to molecular biology data and to automated image analysis system. Currently, we are focused on creating descriptors for the different organs. The ground truth data will be available from March 2011, which will allow us to test our descriptors.
Khalid Niazi, Bettina Selig, Ewert Bengtsson, Ingela Nyström
Partner: Albert Alm, Dept. of Neurosciences, UU
Funding: COMSATS IIT, Islamabad
Period: 0711-
Abstract: Retinal imaging is one of the main sources in ophthalmology to study the optical nerve head and the retina. Retinal images are often used for analyzing, diagnosing and treating a number of diseases of the human retina. Image registration plays an important role in determining the progression of retinal illness. In the current project, we are developing a method which will help in evaluation of glaucoma progression. We are especially concentrating on correction of parallax error, which is normally produced due to a change in the angular position of the camera.
Retinal image registration can be performed between either the full images or within sub-regions. The movement of vessels makes it illogical to perform registration between full images. Using a sub-region which is least effected by vessel movement will present a true picture of the vessel movement. The vessels inside the optic disc, which lie close to the origin, move with time and often get over-exposed during imaging. It has also been reported that the end of the vessel gets detached from the surface of the retina due to loss of the nerve fibers, which leaves us to use the area around the border of the optic disc. We have used the conventional particle swarm optimization (PSO) algorithm which uses uniform distribution to update the velocity equation. Subsequently, we have modified the PSO algorithm which utilizes benefits from the Gaussian and the uniform distribution, when updating the velocity equation. Which one of the distributions is selected depends on the direction o f the cognitive and social components in the velocity equation. This direction checking and selection of the appropriate distribution provide the particles with an ability to jump out of local minima. The registration results achieved by this new version proves the robustness and its ability to find a global minimum.
Our algorithm and results were presented at the SSBA symposium held in Uppsala in March 2010.
Ingrid Carlbom, Milan Gavrilovic, Ewert Bengtsson, Jimmy Azar
Partners: Christer Busch, Marene Landström, Dept. of Genetics and Pathology, UU Hospital
Funding: Swedish Research Council
Period: 1001-
Abstract: Prostate cancer diagnosis is based on Gleason grading, which is the most widely used system for determining the severity of prostate cancer from tissue samples. However, Gleason grading is highly subjective with significant variation between experienced pathologists, which studies show may be as high as 30-40%. We propose to replace subjective diagnosis of prostate cancer with automatic severity grading using a combination of tissue staining and image analysis. New stain combinations enhance color contrast, allowing automatic measurements of morphological features of glands and cells, and thus allowing us to take advantage of computer vision and quantitative image analysis rather than relying on human vision. We intend to demonstrate the accuracy of our severity grading by using the consensus Gleason-graded cohorts from several existing collections of prostate biopsy and prostatectomy material. During 2010, we developed a new staining method particularly suited for image analysis of prostate histopathological data and a technique for color decomposition of these images. By using a combination of histochemical and immunohistochemical staining procedures, we have made it possible to discriminate between normal glandular structures and infiltrating cancer. Thereby we can exclude normal glands from cancer and derive the morphological patterns that characterize malignancy grade progression in these patterns. By using a color model that extracts spectral components of the tissue, we remove the intensity variations that are artifacts from the sectioning and staining procedures. Furthermore, statistical techniques for noise modeling and clustering use soft classification rules allowing linear color decomposition. The purpose is to label each region of the histopathological sections according to its severity grade by employing quantitative image analysis.
Milan Gavrilovic, Carolina Wählby
Partners: Irene Weibrecht, Tim Conze, Ola Söderberg, Ulf Landegren, Dept. of Genetics and Pathology, UU
Funding: TN-faculty, UU
Period: 0807-
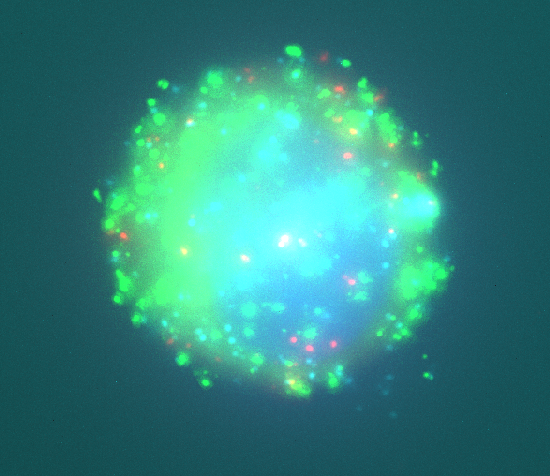
Abstract: Our previously published novel methods for quantification of colocalization are applied to a number of projects in collaboration with the Molecular Tools group at the Dept. of Genetics and Pathology. The aim is detection of multicolour rolling circle products representing DNA-protein interactions in wide-field fluorescence microscopy images. In addition, we use the methods for evaluation of biochemical methods relevant in early stages of specimen preparation, e.g., estimating the quality of oligonucleotides.
Amin Allalou, Carolina Wählby
Partners: Carlos Pardo and Mehmet F. Yanik, Research Laboratory of Electronics, Massachusetts Institute of Technology, Cambridge, USA
Funding: TN-faculty, UU
Period: 1009-
Abstract: During 2010 a research visit was conducted to MIT, Cambridge, USA, and a collaboration was initiated.
A high-throughput platform for cellular resolution in vivo chemical and genetic screens on zebrafish has been developed at MIT, Cambridge, USA. The system automatically loads zebrafish larvae and positions and rotates them for high-speed confocal imaging. The number of neurons in the fish is of importance to different screens and therefore a method has been tested to count the number of neurons in certain regions of the zebrafish brain. In addition, some methods have been developed to rotate and align the fish correctly before imaging.
Amin Allalou, Carolina Wählby
Partners: Nils-Göran Larsson, Karolinska Institute; Mats Nilsson, Chatarina Larsson, Dept. of Genetics and Pathology, UU
Funding: The Swedish Research Council, Collaboration Grant, Medicine
Period: 0801-
Abstract: Mutations of mitochondrial DNA (mtDNA) cause genetic syndromes with widely varying phenotypes and are also implicated in many age-associated diseases and the ageing process itself. Our knowledge of the principles governing segregation of mtDNA mutations in somatic tissues and in the germ line is very limited. In this collaborative project we combine a powerful technique for detection of individual mtDNA molecules with image analysis. We work with a variety of mouse models and the goal is to develop image analysis software to do three-dimensional (3D) reconstruction of the distribution of mutated mtDNA molecules in mammalian tissues. We want to use this technology to study segregation of mtDNA mutations in mouse tissues and to study the mtDNA bottleneck by visualizing the distribution of mutated mtDNA during oogenesis. The ultimate goal is to study the distribution of mtDNA mutations in embryos and placenta to establish principles for prenatal diagnosis.
Amin Allalou, Ida-Maria Sintorn, Carolina Wählby
Partners: Carl-Magnus Clausson, Ola Söderberg, Dept. of Genetics and Pathology, UU
Funding: TN-faculty, UU; VINNMER programme, Swedish Governmental Agency for Innovation Systems
Period: 1002-
Abstract: A novel approach to increase the dynamic range of in situ PLA has been developed at Dept. of Genetics and Pathology, UU. Using several probes with different concentrations the dynamic range can be extended significantly. Signal detection previously developed at CBA, UU, (3DSWD) is used to quantify the number of signals in the different concentrations. See Fig.
Ida-Maria Sintorn
Partners: Adrian Baddeley, Michael Buckley, Leanne Bischof, CSIRO Mathematical and Information Sciences, Australia; Stephen Haggarty, Broad Institute of Harvard and MIT, USA
Funding: S-faculty, SLU; VINNMER programme, Swedish Governmental Agency for Innovation Systems
Period: 0901-
Abstract: In biological research and in the drug development process when screening for new drugs, HCA systems are often used. Such a system is a fully automatized microscopy system that acquires hundreds or thousands of images in an experiment and automatically extracts information from the image data. In this project we develop pre-processing tools to allow for improved comparison of image content within and across cellular screening experiments. During 2010 a method utilizing gradient information for intensity normalization between images was published (Sintorn et al. J. Microscopy). We also develop methods and new statistical analysis tools for so called co-culture HCA experiments i.e., when more than one celltype are cultured together to allow for investigating their interaction in response to added substrates.
Ida-Maria Sintorn
Partner: Vironova AB
Funding: VINNMER programme, Swedish Governmental Agency for Innovation Systems
Period: 0801-
Abstract: Electron Microscopy allows for studying the shape and morphology of biological particles such as viruses at the nm level. This means for example that structural differences between virus maturation stages, related virus species, wild type virus and virus treated with a potential drug or a small molecule can be analyzed. Both external (shape and protein patterns on the virus surface) and internal structural differences can be analyzed. In this project methods for efficiently identifying and quantifying such structural differences are developed.
Gustaf Kylberg, Ida-Maria Sintorn, Ewert Bengtsson, Gunilla Borgefors
Partners: Vironova AB; Ali Mirazimi, Kjell-Olof Höglund, Centre for Microbiological Preparedness, Swedish Institute for Infectious Disease Control (SMI); Jan-Olof Strömberg and Joel Andersson, Dept. of Mathematics, Royal Institute of Technology.
Funding: S-faculty, SLU; Swedish Civil Contingencies Agency; Swedish Defense Materiel Administration; Swedish Governmental Agency for Innovation Systems
Period: 0801-
Abstract: This project is closely linked to Project
In 2010 the project was presented together with Project ![]() at the Swedish National Symposium on Technology and Methodology for Security and Crisis Management (TAMSEC) in Linköping, Sweden.
at the Swedish National Symposium on Technology and Methodology for Security and Crisis Management (TAMSEC) in Linköping, Sweden.

|
Gustaf Kylberg, Ida-Maria Sintorn, Ewert Bengtsson, Gunilla Borgefors
Partners: Vironova AB; Ali Mirazimi, Kjell-Olof Höglund, Centre for Microbiological Preparedness, Swedish Institute for Infectious Disease Control; Gun Frisk and Monika Hodik, Dept. of Immunology, Genetics and Pathology, UU
Funding: S-faculty, SLU; Swedish Civil Contingencies Agency; Swedish Defense Materiel Administration; Swedish Governmental Agency for Innovation Systems
Period: 0801-
Abstract: Transmission electron microscopy (TEM) is an important virus diagnostic tool. The main drawback is that an expert in virus appearance in electron microscopy is needed to perform the analysis at the microscope, an often very time consuming task.
The project aim is to develop methods for a multi-scale analysis at the microscope to automatically acquire highly magnified images of possible virus particles. This is an important step towards automating the virus identification process and thereby creating a rapid, objective, and user independent virus diagnostic system. By introducing the multi-scale approach the search area where highly magnified images are acquired could be decreased by 99.99%. The subsequent segmentation and classification process is developed in Project ![]() . A collaboration has recently been initiated with Dept. of Immunology, Genetics and Pathology, Uppsala University, based on the common interest of automated virus detection using TEM. The project was presented together with Project
. A collaboration has recently been initiated with Dept. of Immunology, Genetics and Pathology, Uppsala University, based on the common interest of automated virus detection using TEM. The project was presented together with Project ![]() at the Swedish National Symposium on Technology and Methodology for Security and Crisis Management (TAMSEC) in Linköping, Sweden.
at the Swedish National Symposium on Technology and Methodology for Security and Crisis Management (TAMSEC) in Linköping, Sweden.
Azadeh Fakhrzadeh, Cris Luengo, Gunilla Borgefors
Partners: Ellinor Spörndly-Nees, Lena Holm, Dept. of Anatomy, Physiology and Biochemistry, SLU
Funding: SLU (KoN)
Period: : 1009-
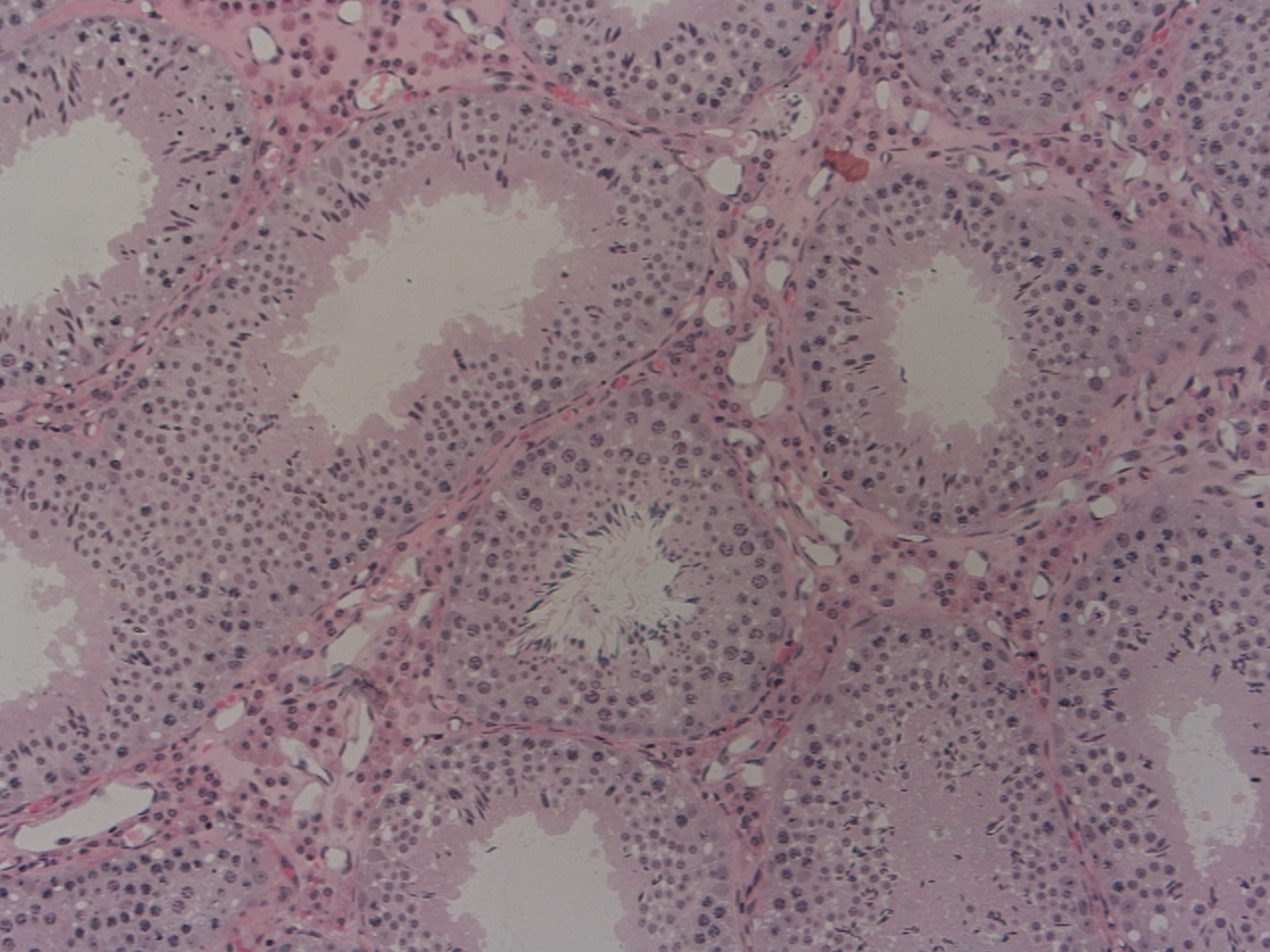
Abstract: Reproductive toxicology is the study of chemicals and their effects on the reproductive system of human and animal. In reproductive toxicology, there is a strong need to detect structural differences in organs that often have both a complex microscopic structure and function. This problem is further complicated because standard techniques are based on the examination of two-dimensional sections of a three-dimensional structure. The aim of this project is to develop methods to objectively describe microscopic structures of male reproductive organs and to test these in reproductive toxicology research. The project is comparative and may include studies of organs from fish, frog and birds as well as wild and domesticated mammals. We will develop automatic and interactive methods to analyze the relevant structures in the histology images of testis, see Fig.
Hamid Sarve, Joakim Lindblad, Vladimir Curic, Gunilla Borgefors
Partners: Carina Johansson, Dept. of Clinical Medicine, Örebro University; Nataša Sladoje, Faculty of Technical Sciences, University of Novi Sad, Serbia
Funding: Swedish Research Council; S-faculty, SLU
Period: 0503-
Abstract: With an aging and increasingly osteoporotic population, bone implants are becoming more important to ensure the quality of life. In order to evaluate how tissue reacts on implants, the interface at the implant and tissue must be studied. Today, this is done manually in a microscope which is a costly and time-consuming procedure.
The aim of this project is to develop automatic image analysis methods for evaluating images of the interface region of tissue and implant. These methods would provide faster and more objective measurements on how well the implant is integrated in the bone compared to today's manual methods. The analysis involves segmentation of the images into different tissue-types and quantifying bone contact length, area and volume.
The project encompasses parallel development and comparison of methods for 2D analysis of histological sections as well as 3D analysis of SRCT volumes. Within the project, methods for segmentation and feature extraction have been developed for both 2D histological sections, and 3D SRCT volumes. To facilitate comparison of results from the two imaging modalities, a 2D-3D image registration method has also been developed. During 2010, by extending the features traditionally used for 2D analysis, methods for extraction of the 3D features from the 3D volume data were developed. This work was accepted for publication in the Journal of Computer Methods and Programs in Biomedicine.
Hamid Sarve, Joakim Lindblad, Gunilla Borgefors
Partners: Carina Johansson, Dept. of Clinical Medicine, Örebro University; AstraTech, Mölndal
Funding: The Knowledge Foundation
Period: 0906-
Abstract: This project aims to develop new techniques for interactive 3D visualization of bone anchored implants in order to facilitate the understanding of the mechanisms of implant integration. To enable good communication between the people involved in development, production and use of medical devices - computer scientists, material scientists, and medical doctors - each with their own special knowledge, it is of highest importance to provide a common visual platform for a mutual understanding of the problems of implant integration. Being able to actually see the 3D structure around the implant for the first time will inspire new measures of the implant integration quality.
We base the visualization on data from non-destructive 3D SRCT imaging; this technique yields more accurate tomographic reconstruction at higher resolution compared to standard CT. Furthermore, SRCT-imaging is more suitable for samples containing metal, as the artefacts caused by the metal is significantly lower compared to traditional CT. Existing visualization software are not really useful for this type of complex and highly detailed data, requiring the development of special purpose methods and software.
The combination of this project and Project ![]() will improve both quantitative and qualitative analysis of bone implant integration and thereby support the development of more effective implants and diminish the number of malfunctioning devices. During 2010, two new methods for visualizing SRCT-scanned volume samples were presented at the International Conference on Computer Vision and Graphics (ICCVG'10); one being an animation that follows the implant thread and extracts information about the bone-implant integration over the whole sample and the other a 2D unfolding that displays a flattened version of the implant surface, with feature information projected onto it, providing a direct overview of the implant integration (see Fig.
will improve both quantitative and qualitative analysis of bone implant integration and thereby support the development of more effective implants and diminish the number of malfunctioning devices. During 2010, two new methods for visualizing SRCT-scanned volume samples were presented at the International Conference on Computer Vision and Graphics (ICCVG'10); one being an animation that follows the implant thread and extracts information about the bone-implant integration over the whole sample and the other a 2D unfolding that displays a flattened version of the implant surface, with feature information projected onto it, providing a direct overview of the implant integration (see Fig. ![]() ).
).
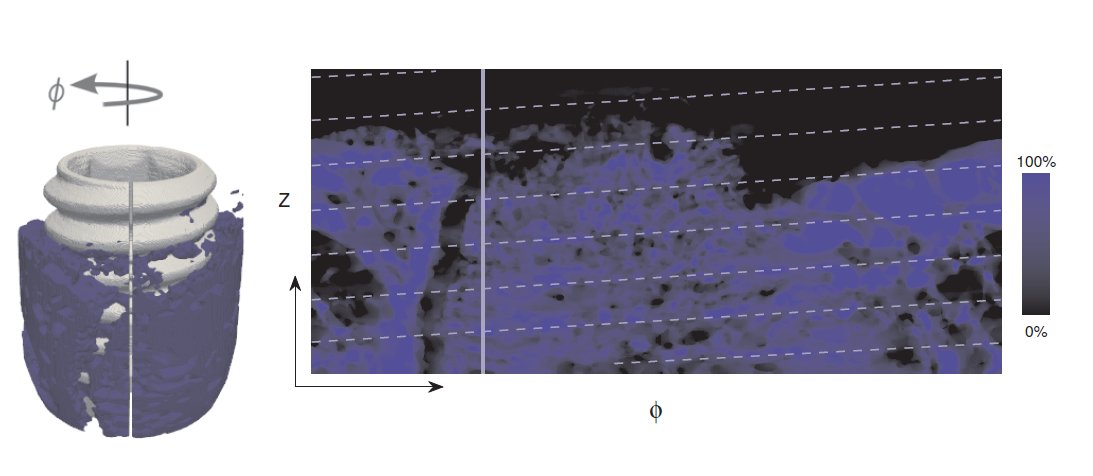
|
Stina Svensson, Magnus Gedda
Partner: Dept. of Cell and Molecular Biology, Karolinska Institutet, Stockholm
Funding: Swedish Research Council (Project 621-2005-5540); SLU S-faculty; UU TN-faculty
Period: 0401-1005
Abstract: State of the art imaging techniques makes it possible to study individual proteins and other macromolecules from a structural point of view. Descriptions with respect to geometry and shape facilitates studying protein dynamics. This type of study is essential to increase the understanding of their biological role. CMB has developed methods, using cryo electron tomography (Cryo-ET), for 3D imaging of individual proteins at a resolution of a few nm. Localisation of the protein in the image has so far been done manually. For large-scale studies, computerized image analysis serves as an essential tool to automatically and objectively analyse image content. In this project, we develop methods to automatically localise macromolecules from in vitro Cryo-ET images and thereby make large-scale studies of proteins possible. This is done by taking into account both grey-level information (which reflects the internal structure of the protein) and 3D shape information.
Andreas Kårsnäs, Robin Strand, Ewert Bengtsson, Stefan Seipel
Partners: Visiopharm, Hörsholm, Denmark; Clinical Pathology Division, Vejle hospital, Vejle, Denmark
Funding: NordForsk Private Public Partnership PhD Programme and Visiopharm
Period: 0909-
Abstract: The results of analyses of tissue biopsies by pathologists are crucial for breast cancer patients. In particular, the precision of a patient's prognosis, and the ability to predict the consequences of various treatment opportunities before actually exposing the cancer patient, depend on the detection and quantification of biomarkers in tissue sections by microscopy. Experience from the last decade has revealed that manual detection and quantification of biomarkers by microscopy of tissue biopsies is highly dependent on the competencies and stamina of the individual pathologist. The aim of the present PhD project is to develop software-based algorithms, which can facilitate the workflow and ensure objective and more precise results of the quantitative microscopy procedures in breast cancer. During spring 2010, we finished the first part of the development of the Pathology Module, a user interface supporting pathologists while ensuring objective and more precise results of the quantitative microscopy procedures in breast cancer. During the fall 2010, we have started a project for automatic detection of cancer cells. The main focus has been on detecting cell nuclei, and we have, in collaboration with DTU in Denmark, developed an algorithm for nuclei-detection and separation of co-localized nuclei.